Brief Review — PubMedQA: A Dataset for Biomedical Research Question Answering
PubMedQA: Yes/No/Maybe QA Dataset

PubMedQA: A Dataset for Biomedical Research Question Answering
PubMedQA, by University of Pittsburgh, Carnegie Mellon University, Google AI,
2019 EMNLP-IJCNLP, Over 230 Citations (Sik-Ho Tsang @ Medium)Medical NLP
==== My Other Paper Readings Are Also Over Here ====
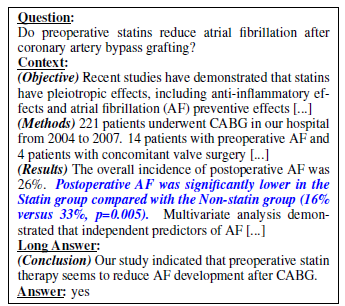
- PubMedQA, a novel biomedical question answering (QA) dataset, is introduced, which is collected from PubMed abstracts.
- The task of PubMedQA is to answer research questions with yes/no/maybe using the corresponding abstracts.
- PubMedQA has 1k expert-annotated, 61.2k unlabeled and 211.3k artificially generated QA instances.
- (This is a dataset evaluated by Med-PaLM.)
Outline
- PubMedQA
- BioBERT Fine-Tuning
- Results
1. PubMedQA
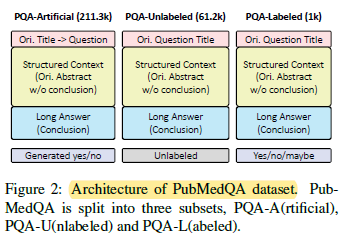
- PubMedQA is split into 3 subsets: labeled, unlabeled and artificially generated. They are denoted as PQA-L(abeled), PQA-U(nlabeled) and PQA-A(rtificial), respectively, as above.
- Collection of PQA-L and PQA-U: PubMed articles which have i) a question mark in the titles and ii) a structured abstract with conclusive part are collected and denoted as pre-PQA-U.
1.1. PQA-L
- Two annotators labeled 1k instances from pre-PQA-U with yes/no/maybe to build PQA-L.
- Reasoning-free setting: The annotator 1 doesn’t need to do much reasoning to annotate since the long answer is available.
- Reasoning-required setting: Annotator 2 cannot use the long answer, so reasoning over the context is required for annotation.
1.2. PQA-U
- The unlabeled instances in pre-PQA-U with yes/no/maybe answerable questions are included to build PQA-U.
1.3. PQA-A

- A simple heuristic is used to collect many noisily-labeled instances to build PQA-A for pretraining.
- 2 examples are shown above.
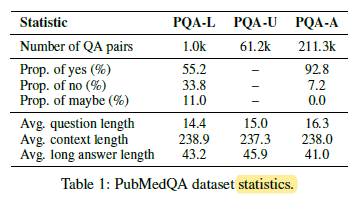
1.4. PubMedQA Statistics

- Figure 3: PubMed abstracts are manually annotated by medical librarians with Medical Subject Headings (MeSH). MeSH terms are used to represent abstract topics, and their distribution is visualized.
- Most (57.5%) involve comparing multiple groups (e.g.: experiment and control), and others require interpreting statistics of a single group or its subgroups. Reasoning over quantitative contents is required in nearly all (96.5%) of them.
- Figure 4: The Sankey diagram is used to show the proportional relationships.

2. BioBERT Fine-Tuning
2.1. Loss Function
- With regularizing neural machine translation models with binary bag-of-word (BoW) statistics, BioBERT is fine-tuned with an auxiliary task of predicting the binary BoW statistics of the long answers:
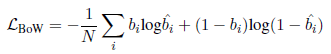
- where bi and ^bi are ground-truth and predicted probability of whether token i is in the long answers. N is the BoW vocabulary size.
- The total loss is:
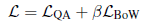
2.2. Multi-Phase Fine-Tuning Architecture
Conceptually, two BioBERTs are trained. One is trained to generate pseudo-labels for PQA-U, which can enlarge the training set. Another one is trained using both labeled and pseudo-labelled dataset for later evaluation.
- (Please skip the details below for quick reading.)
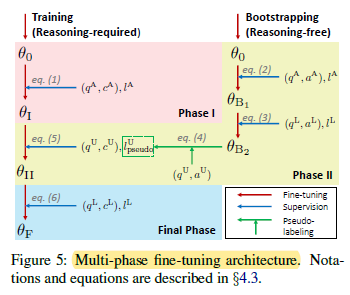
- q, c, a, l are used to denote question, context, long answer and yes/no/maybe label of instances, respectively.
2.2.1. Phase I Fine-tuning on PQA-A
- Pretrained BioBERT is initialized with θ0, and fine-tuned on PQA-A using question and context as input:
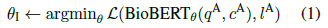
2.2.2. Phase II Fine-tuning on Bootstrapped PQA-U
- To fully utilize the unlabeled instances in PQA-U, pseudo-labeling with a bootstrapping is strategy used. BioBERT is initialized with θ0, and fine-tuned on PQA-A using question and long answer (reasoning-free):

- Then further fine-tune BioBERTθB1 on PQA-L:
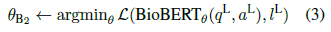
- Pseudo-label PQA-U instances using the most confident predictions of BioBERTθB2 for each class, are used:

- Next, BioBERTθI is fine-tuned on the bootstrapped PQA-U using question and context (under reasoning-required setting):
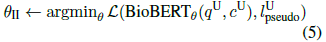
2.2.3. Final Phase Fine-tuning on PQA-L
- In the final phase, BioBERTθII is fine-tuned on PQA-L:
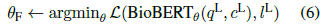
- Final predictions on instances of PQA-L validation and test sets are made using BioBERTθF:
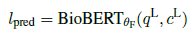
3. Results
3.1. Human Baseline
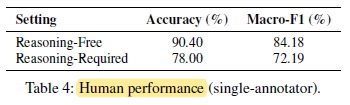
3.2. PQA-L Test Set
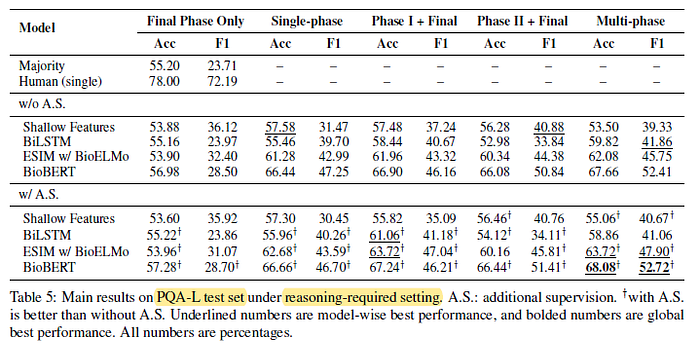
In general, multi-phase fine-tuning of BioBERT with additional supervision outperforms other baselines by large margins, but the results are still much worse than just single-human performance.
- (Please read the paper directly for more experimental results.)
