Brief Review — An Efficient Solution for Breast Tumor Segmentation and Classification in Ultrasound Images Using Deep Adversarial Learning
Conditional GAN (cGAN) + Atrous Convolution (AC) + Channel Attention with Weighting Block (CAW)
An Efficient Solution for Breast Tumor Segmentation and Classification in Ultrasound Images Using Deep Adversarial Learning
cGAN+AC+CAW, by Universitat Rovira i Virgili, and Bioinformatics Institute,
2019 arXiv v1 (Sik-Ho Tsang @ Medium)
Medical Image Analysis, Image Segmentation, Image Classification
- An efficient solution for tumor segmentation and classification in breast ultrasound (BUS) images. An atrous convolution layer is added to the conditional generative adversarial network (cGAN) segmentation model. A channel-wise weighting block is also added.
- SSIM and L1-norm loss with the typical adversarial loss are proposed as a loss function.
1. cGAN+AC+CAW for Segmentation

1.1. Generator G
- The generator network incorporates an encoder section, made of seven convolutional layers (En1 to En7), and a decoder section, made of seven deconvolutional layers (Dn1 to Dn7).
- An atrous convolution block is inserted between En3 and En4. 1, 6 and 9 dilation rates with kernel size 3×3 and a stride of 2 are used.
- in addition to a channel attention with channel weighting (CAW) block between En7 and Dn1.
- The CAW block is an aggregation of a channel attention module (DAN) with channel weighting block (SENet), which increases the representational power of the highest level features of the generator network.
1.2. Discriminator D
- It is a sequence of convolutional layers.
- The input of the discriminator is the concatenation of the BUS image and a binary mask marking the tumor area.
- The output of the discriminator is a 10×10 matrix having values varying from 0.0 (completely fake) to 1.0 (real).
1.3. Loss Function
- The loss function of the generator G comprises three terms: adversarial loss (binary cross entropy loss), L1-norm to boost the learning process, and SSIM loss to improve the shape of the boundaries of segmented masks:
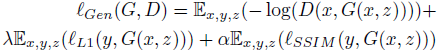
- where z is a random variable.
- The loss function of the discriminator D is:

2. Random Forest for Classification
- Each BUS image is fed into the trained generative network to obtain the boundary of the tumor, and then 13 statistical features from that boundary are computed: fractal dimension, lacunarity, convex hull, convexity, circularity, area, perimeter, centroid, minor and major axis length, smoothness, Hu moments (6) and central moments (order 3 and below).
- Exhaustive Feature Selection (EFS) algorithm is used to select the best set of features. The EFS algorithm indicates that the fractal dimension, lacunarity, convex hull, and centroid are the 4 optimal features.
- The selected features are fed into a Random Forest classifier, which is later trained to discriminate between benign and malignant tumors.
3. Results
3.1. Segmentation
- This dataset contains 150 malignant and 100 benign tumors contained in BUS images. To train our model, we randomly divided the dataset into the training set (70%), a validation set (10%) and testing set (20%).
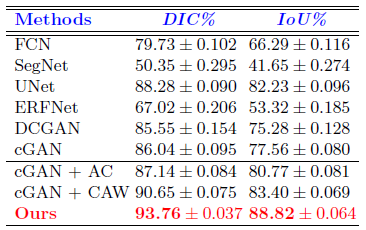
The model (cGAN+AC+CAW) outperforms the rest in all metrics. It achieves Dice and IoU scores of 93.76% and 88.82%, respectively.
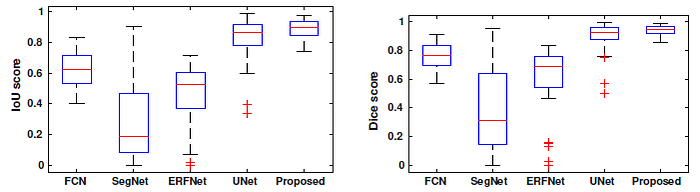
The proposed model is in the range 88% to 94% for Dice coefficient and 80% to 89% for IoU, while other deep segmentation methods, FCN, SegNet, ERFNet and U-Net show a wider range of values.

- SegNet and ERFNet yield the worst results since there are large false negative areas (in red), as well as some false positive areas (in green).
In turn, U-Net, DCGAN, cGAN provide good segmentation but the proposed model provide more accurate segmentation of the boundary of breast tumors.
3.2. Classification

- The proposed breast tumor classification method outperforms [9], with a total accuracy degree of 85%.
Reference
[2019 arXiv v1] [cGAN+AC+CAW]
An Efficient Solution for Breast Tumor Segmentation and Classification in Ultrasound Images Using Deep Adversarial Learning
Biomedical Multi-Task Learning
2018 [ResNet+Mask R-CNN] [cU-Net+PE] [Multi-Task Deep U-Net] [cGAN-AutoEnc & cGAN-Unet] 2019 [cGAN+AC+CAW]
