Review — Self-supervised Feature Learning for 3D Medical Images by Playing a Rubik’s Cube
Rubik’s Cube Recovery: Cube Rearrangement and Cube Rotation
Self-supervised Feature Learning for 3D Medical Images by Playing a Rubik’s Cube, Rubik’s Cube, by Tsinghua University, and Tencent,
2019 MICCAI, Over 70 Citations (Sik-Ho Tsang @ Medium)
Self-Supervised Learning, Image Classification, Medical Image Classification, Medical Image Segmentation
- A self-supervised learning framework is proposed for the volumetric medical images, which is the first work focusing on the self-supervised learning of 3D neural networks.
- A proxy task, i.e., Rubik’s cube recovery, is formulated to pretrain 3D neural networks. The task involves two operations, i.e., cube rearrangement and cube rotation, which enforce networks to learn translational and rotational invariant features.
- Finally, the pretrained network is fine-tuned for brain hemorrhage classification and brain tumor segmentation.
Outline
- Rubik’s Cube Recovery
- Experimental Results
1. Rubik’s Cube
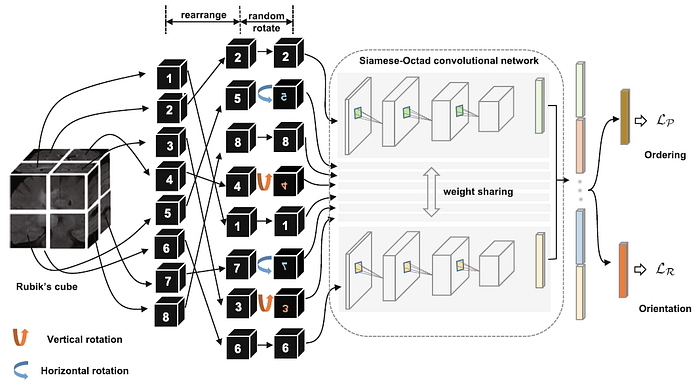
- A 3D medical volume is first partitioned it into a grid (e.g., 2×2×2) of cubes, and then permuted with random rotations. Like playing a Rubik’s cube, the proxy task aims to recover the original configuration, i.e., cubes are ordered and orientated.
1.1. Preprocessing
- To encourage the network to learn and use high-level semantic features for Rubik’s cube recovery, rather using the texture information that close to the cube boundaries, a gap (about 10 voxels) between two adjacent cubes is used.
- The cube intensities are normalized to [−1, 1].
1.2. Network Architecture
- A Siamese network with M (which is the number of cubes) sharing weight branches, namely Siamese-Octad, is adopted to solve Rubik’s cube.
- 3D CNN, e.g., 3D VGG is used.
- The feature maps from the last fully-connected or convolution layer of all branches are concatenated and given as input to the fully-connected layer of separate tasks, i.e., cube ordering and orientating, which are supervised by permutation loss (LP) and rotation loss (LR), respectively.
1.3. Cube Ordering
- For each time of Rubik’s cube recovery, the eight cubes are rearranged according to one of the K permutations, e.g., (2, 5, 8, 4, 1, 7, 3, 6) in the above figure.
- The network is trained to identify the selected permutation from the K options, which can be seen as a classification task with K categories.
- Assuming the 1×K network prediction as p and the one-hot label as l, the permutation loss (LP) in this step can be defined as:

1.4. Cube Orientation
- To reduce the complexity of the task, the directions for cube rotation are limited, i.e., only allowing 180◦ horizontal and vertical rotations.
- As shown above, the cubes (5, 7) and (4, 3) are horizontally and vertically rotated, respectively.
- The network is required to recognize whether each of the input cubes has been rotated. It can be seen as a multi-label classification task using the 1×M (M is the number of cubes) ground truth (g) with 1 on the positions of rotated cubes and 0 vice versa.
- The predictions of this task are two 1×M vectors (r) indicating the possibilities of horizontal (hor) and vertical (ver) rotations for each cube.
- The rotation loss (LR) is:
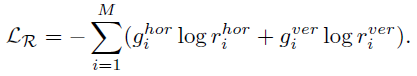
1.5. Full Objective
- The full objective (L) is:
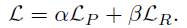
- where equal weights of α=β=0.5 leads to the best feature representations.
1.6. Target Tasks
- For the classification task, the pre-trained CNN can be directly used for fine-tuning.
- For the segmentation of 3D medical images, the pre-trained weights can only be adapted to the encoder part of the fully convolutional network (FCN), e.g. U-Net. The decoder of FCN still needs random initialization, which may wreck the pre-trained feature representation and neutralize the improvement generated by the pre-training.
- Inspired by the Dense Upsampling Convolution (DUC) in ResNet-DUC-HDC, convolutional operations are applied directly on feature maps yield by the pre-trained encoder to get the dense pixel-wise prediction instead of the transposed convolutions. The DUC can significantly decrease the number of trainable parameters of the decoder and alleviate the influence caused by random initialization.
2. Experimental Results
2.1. Datasets
2.1.1. Brain Hemorrhage Dataset
- 1486 brain CT scan images are collected from a collaborative hospital. The 3D CT volumes containing brain hemorrhage can be classified to four pathological causes, i.e., aneurysm, arteriovenous malformation, moyamoya disease and hypertension. Each 3D CT volume is of size 230×270×30 voxels.
- The cube size of Rubik’s cube is 64×64×12.
- The average classification accuracy (ACC) is adopted as metric.
2.1.2. BraTS-2018 Dataset
- The training set consists of 285 brain tumor MR volumes, which have four modalities, i.e., native T1-weighted (T1), post-contrast T1-weighted (T1Gd), T2-weighted (T2), and T2 Fluid Attenuated Inversion Recovery (FLAIR). The size of each volume is 240×240×155 voxels.
- The cube size of Rubik’s cube is 64×64×64. As the BraTS-2018 has four modalities, the cubes from different modalities are concatenated and each branch of Siamese-Octad network is sent as input.
- The mean intersection over union (mIoU) is adopted as the metric to evaluate the segmentation accuracy.
2.2. Pretraining Accuracy

- As the random cube rotation increases the difficulty of solving Rubik’s cube, the test accuracies of cube ordering degrade with −7.7% and −6.6% for brain hemorrhage dataset and BraTS-2018, respectively.
- On the other hand, the Rubik’s cube network can achieve test accuracies of 93.1% and 82.1% for the cube orientation.
2.3. Performance of Target Tasks
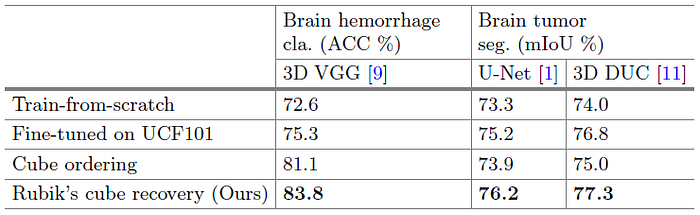
- The train-from-scratch strategy is involved as the baseline.
- For UCF-101 pretrained models, frames are extracted from UCF-101 videos to form a cube of 112×112×16 to pre-train the 3D network.
- The proposed Rubik’s cube pre-trained weights are generated by deeply exploiting useful information from limited training data without using any extra dataset.
- On Brain Hemorrhage Dataset, due to the gap between natural video and volumetric medical data, the improvement yielded by UCF101 pre-trained weights is limited, i.e., +2.7%. In comparison, the proposed Rubik’s cube pretrained weights substantially boost the classification accuracy to 83.8%, which is 11.2% higher than that of train-from-scratch model.
- On Brain Tumor dataset, the model fine-tuned from our Rubik’s cube recovery paradigm can generate more accurate segmentations for brain tumors, i.e., mIoUs of 76.2% and 77.3% are achieved by the U-Net and 3D DUC, respectively.
Inspired from Jigsaw, Rubik’s cube pretext task is designed for 3D image self-supervised learning.
Reference
[2019 MICCAI] [Rubik’s Cube]
Self-supervised Feature Learning for 3D Medical Images by Playing a Rubik’s Cube
Self-Supervised Learning
1993 … 2019 [Rubik’s Cube] … 2021 [MoCo v3] [SimSiam] [DINO] [Exemplar-v1, Exemplar-v2] [MICLe] [Barlow Twins]
Biomedical Image Classification
2019 [CheXpert] [Rubik’s Cube] 2020 [VGGNet for COVID-19] [Dermatology] 2021 [MICLe]
Biomedical Image Segmentation
2015 … 2019 [DUNet] [NN-Fit] [Rubik’s Cube] 2020 [MultiResUNet] [UNet 3+] [Dense-Gated U-Net (DGNet)]