Swin-SFTNet: Spatial Feature Expansion and Aggregation Using Swin Transformer For Whole Breast Micro-Mass Segmentation
Swin-SFTNet, Swin Transformer as Backbone, SFEA for Better Skip Connection
Swin-SFTNet: Spatial Feature Expansion and Aggregation Using Swin Transformer For Whole Breast Micro-Mass Segmentation,
SWIN-SFTNet, by University of Nevada,
2022 arXiv v1 (Seems 2023 ISBI) (Sik-Ho Tsang @ Medium)Biomedical Image Segmentation
2015 … 2022 [UNETR] [Half-UNet] [BUSIS] [RCA-IUNet] [Swin-Unet] [DS-TransUNet] [UNeXt] [AdwU-Net] [TransUNetV2] 2023 [DCSAU-Net] [RMMLP]
==== My Other Paper Readings Are Also Over Here ====
- Swin-SFTNet, a novel U-Net-shaped Transformer-based architecture, is proposed.
- To capture the global context, a novel Spatial Feature Expansion and Aggregation Block (SFEA) is designed that transforms sequential linear patches into a structured spatial feature.
- A novel embedding loss is incorporated that calculates similarities between linear feature embeddings of the encoder and decoder blocks.
Outline
- Swin-SFTNet
- Loss Function
- Results
1. Swin-SFTNet
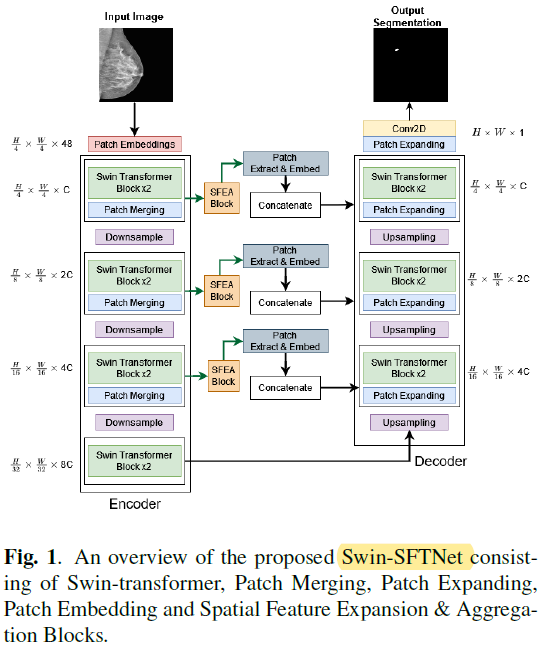
1.1. Input
- The breast mammography grayscale images are firstly transformed into RGB (maybe increase the channel size only?), providing the model with learning essential features.
- A patch-embedding layer is utilized to transform the input into non-overlapping patches of size 4×4. So for three RGB channels, we get to 4×4×3=48 depth dimension.
1.2. Encoder
- The encoder blocks, each consisting of two successive Swin Transformer blocks and a patch-merging layer.
- The Swin Transformer block is:

- where Φ is Layer Norm layer, δ is GELU.
- The encoder blocks are repeated three times to downsample the feature dimensions.
1.3. Decoder
- A symmetric decoder is used, which is composed of multiple Swin Transformer blocks and patch expanding layer.
- Each decoder block is concatenated with the skip-connection features from the encoder with the same spatial dimension.
- Decoder blocks are repeated for three times to upsample.
1.4. Output
- The last patch-expanding layer is incorporated to perform 4× up-sampling to restore the resolution.
- At last, a 2D convolution is applied to get the output feature dimension H×W×1 for binary mass segmentation.
- Here, H=256, W=256 and C=128.
1.5. Spatial Feature Expansion and Aggregation (SFEA)
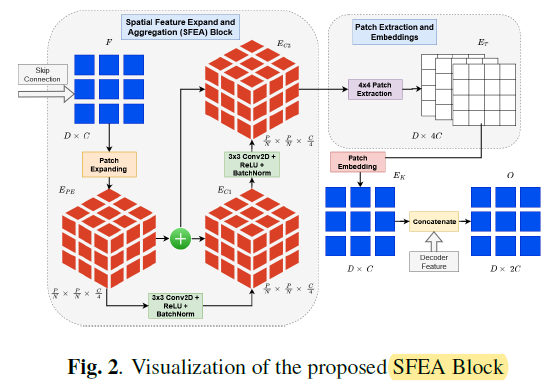
- A patch expanding layer with patch-size 4×4 which gives out the feature output EPE.
- Next, a 3×3 2D Convolution, ReLU activation, and Batch-Normalization operation followed by element-wise addition of features are applied to EPE to get output feature EC1.
- In a similar manner, another same 2D Convolution Block is applied on EC1 to get feature output and element-wise features are added from EPE to get final output EC2.
These two convolution operations help with extracting global spatial context information that is further combined with the decoder’s local patch-level information.
- 4×4 Patch-extraction operation is used to convert the feature back into 2D sequence feature output, ET.
- After that, patch-embedding layer is used to make the feature dimension same as the decoder’s paired output, so the output feature map becomes EK.
- Next, the feature from the decoder’s patch expanding layer is concatenated with the EK.
- Three different values for C=[128, 256, 512] are used for the three skip connections.
2. Loss Function
- Binary cross-entropy loss is used:
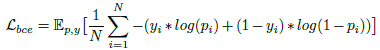
- Dice loss is also used:
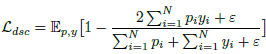
- A novel embedding loss is also used by obtaining positional and patch features from the Transformer encoder layers E and decoder layers D.

- where Q stands for the number of features extracted from the embedding layers of the Transformer encoder.
- Finally, the total loss is:
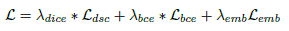
- where λbce=0.4, λdice=0.6 and λemb=0.01.
3. Results
3.1. Qualitative Results

- The model can segment harder and smaller masses than other architectures.
3.2. SOTA Comparisons
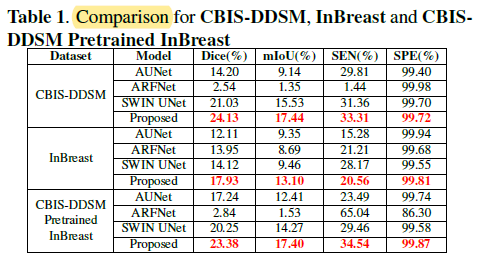
The proposed model achieves the best score compared to others for all the metrics.
3.3. Ablation Studies

With the novel loss function, 3.14%, 6.33%, and 0.86% gains are obtained for CBISDDSM, InBreast, and CBIS pre-trained model consecutively.
