Review — Big Bird: Transformers for Longer Sequences
BIGBIRD, Turing Complete, Sparse Attention Approximates Full Attention
Big Bird: Transformers for Longer Sequences,
BIGBIRD, by Google Research,
2020 NeurIPS, Over 900 Citations (Sik-Ho Tsang @ Medium)
NLP, Transformer, Longformer
==== My Other Paper Readings Also Over Here ====
- BIGBIRD, a sparse attention mechanism that reduces the Transformer quadratic dependency to linear.
- The proposed sparse attention can handle sequences of length up to 8× of what was previously possible using similar hardware.
Outline
- Generalized Attention Mechanism & Full Attention in Transformer
- BIGBIRD
- NLP Results
- Genomics Results
1. Generalized Attention Mechanism & Full Attention in Transformer
1.1. Generalized Attention Mechanism
- On an input sequence X=(x1, …, xn) of size n×d, the generalized attention mechanism is described by a directed graph D whose vertex set is [n] = {1, …, n}.
- The set of arcs (directed edges) represent the set of inner products that the attention mechanism will consider.
- Let N(i) denote the out-neighbors set of node i in D, then the i-th output vector of the generalized attention mechanism is defined as:
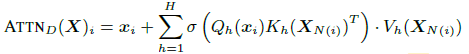
- where Qh, Kh, and Vh are query, key, and value respectively. σ is scoring function, i.e. softmax. H is the number of heads.
- XN(i) corresponds to the matrix formed by only stacking {xj: j ∈ N(i)} and NOT all the inputs.
1.2. Full Attention in Transformer
If D is the complete digraph, we recover the full quadratic attention mechanism in the conventional Transformer.
- To elaborate, A∈[0, 1] of size n×n with A(i, j)=1 if query i attends to key j and is zero otherwise.
However, full attention induces quadratic complexity.
1.3. Graph Sparsification Problem
The problem of reducing the quadratic complexity of self-attention can now be seen as a graph sparsification problem.
- It is well-known that random graphs are expanders and can approximate complete graphs in a number of different contexts including in their spectral properties [80, 38].
2. BIGBIRD
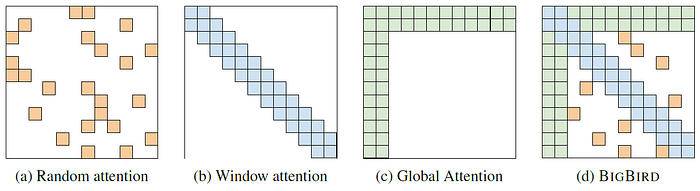
2.1. (a) Random Attention
- Let us consider the simplest random graph construction, known as Erdős–Rényi model, where each edge is independently chosen with a fixed probability.
A sparse attention where each query attends over r random number of keys, i.e. A(i, ·)=1 for r randomly chosen keys.
2.2. (b) Sliding Window Attention
- The second viewpoint which inspired the creation of BIGBIRD is that most contexts within NLP and computational biology have data which displays a great deal of locality of reference.
- A particular model introduced by Watts and Strogatz [95] is of high relevance to BIGBIRD as it achieves a good balance between average shortest path and the notion of locality.
The generative process of their model is as follows: Construct a regular ring lattice, a graph with n nodes each connected to w neighbors, w/2 on each side, to capture the local structures in the context.
- Then, we can have a random subset (k%) of all connections is replaced with a random connection. The other (100 — k)%, local connections are retained.
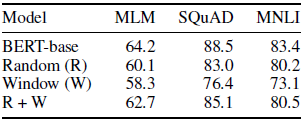
- When random (R) and sliding window (W) are used together for attention, the results are closed to BERT one.
2.3. (c) Global Attention
- The final piece is the importance of “global tokens” (tokens that attend to all tokens in the sequence and to whom all tokens attend to.)
BIGBIRD-ITC: Internal Transformer Construction (ITC) make some (g) existing tokens “global”.
BIGBIRD-ETC: Extended Transformer Construction (ETC) includes g additional “global” tokens such as CLS.
2.4. (d) BIGBIRD Attention
The final attention mechanism for BIGBIRD has all three of these properties: queries attend to r random keys, each query attends to w/2 tokens to the left of its location and w/2 to the right of its location and they contain g global tokens (The global tokens can be from existing tokens or extra added tokens).
- Authors also show the full attention is somehow Turing Complete.
- BIGBIRD is a universal approximator of sequence functions and is Turing Complete, thereby preserving these properties of the quadratic, full attention model.
- (Please feel free to read the paper for more details.)
3. NLP Results
3.1. QA Tasks

- BIGBIRD-ETC, with expanded global tokens consistently outperforms all other models. Thus, this configuration is chosen to train a large sized model to be used for evaluation on the hidden test set.
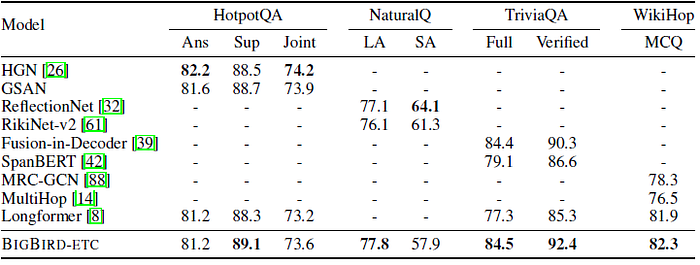
- BIGBIRD-ETC model is compared to top-3 entries from the leaderboard excluding BIGBIRD.
- It is worth noting that BIGBIRD submission is a single model, whereas the other top-3 entries for Natural Questions are ensembles.
One can clearly see the importance of using longer context as both Longformer and BIGBIRD outperform models with smaller contexts.
3.2. Summarization

- Following BERT, one layer with cross entropy loss is used on top of the first [CLS] token.
Gains of using BIGBIRD are more significant when longer documents and fewer training examples are used.
4. Genomics Results
- There has been a recent upsurge in using deep learning for genomics data [87, 107, 13], which has resulted in improved performance on several biologically-significant tasks such as promoter site prediction [71], methylation analysis [55], predicting functional effects of non-coding variant [110], etc.
4.1. Bits Per Character (BPC)

- DNA is first segmented into tokens so as to further increase the context length. In particular, a byte-pair encoding table is built for the DNA sequence of size 32K, with each token representing 8.78 base pairs on average. The the bits per character (BPC) on a held-out set is shown above.
Attention based contextual representation of DNA does improve BPC, which is further improved by using longer context.
4.2. Promoter Region Prediction
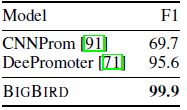
- Promoter is a DNA region typically located upstream of the gene, which is the site of transcription initiation.
BIGBIRD achieve nearly perfect accuracy with a 5% jump from the previous best reported accuracy.
4.3. Chromatin-Profile Prediction

- Non-coding regions of DNA do not code for proteins. Majority of diseases and other trait associated single-nucleotide polymorphism are correlated to non-coding genomic variations [110, 46].
- The corresponding ML task is to predict, for a given non-coding region of DNA, these 919 chromatin-profile including 690 transcription factors (TF) binding profiles for 160 different TFs, 125 DNase I sensitivity (DHS) profiles and 104 histone-mark (HM) profiles. 919 binary classifiers are jointly learned to predict these functional effects from sequence of DNA fragments.
- On held-out chromosomes, AUC is measured and compared with the baselines.
BIGBIRD significantly improves on performance on the harder task HM, which is known to have longer-range correlations [27] than others.
Reference
[2020 NeurIPS] [BIGBIRD]
Big Bird: Transformers for Longer Sequences
2.1. Language Model / Sequence Model
1991 … 2020 … [BIGBIRD] 2021 [Performer] [gMLP] [Roformer] [PPBERT] [DeBERTa] [DeLighT] [Transformer-LS] 2022 [GPT-NeoX-20B] [InstructGPT]
